- Home
- Species & Infections
- Candida Guilliermondii
Candida guilliermondii - A Rare Fungal Pathogen
Updated 6/15/2025
Written by Molecular Biologist Dr. Vibhuti Rana, PhD
Very little is known about a non Candida albicans yeast species called Candida guilliermondii, that can invade host bodies and cause a myriad of fungal infections. This article will describe all relevant information you need to know about C. guilliermondii.
While majority of the fungal infections are caused by Candida albicans, the non-albicans yeast species of the genus Candida like C. tropicalis, C. parapsilosis, and C. glabrata are answerable for 35–65% of all candidemia cases in patients. While some of the fungal infections may not cause much trouble, most of them can get serious beyond control if not tackled effectively.
C. guilliermondii gained clinical relevance when it started showing multi-drug resistance to popular antimycotic agents (1). It is known (in reality, misidentified due to mixed characteristics with other similar species) by some other names like Pichia guilliermondii, C. famata, Candida fermentati (Pichia caribbica), and Candida carpophila. Thanks to the molecular assays like electrophoretic karyotyping, RAPD (random amplified polymorphic DNA) analysis, study of chromosomal DNA binding profiles, as well as of DNA composition, scientists were able to differentiate between the highly confusing Pichia guilliermondii and Pichia caribbica and identify them as separate species (1).
Ultimately, in all cases of uncertainties, genomic analysis works best than any of the microscopic or biochemical tests for accurate identification of this pathogenic yeast species. As of now, the C. guilliermondii complex comprises C. guilliermondii sensu stricto, C. fermentati, C. carpophila, and other species (2).
C.
guilliermondii's Phenotype Switching Abilities
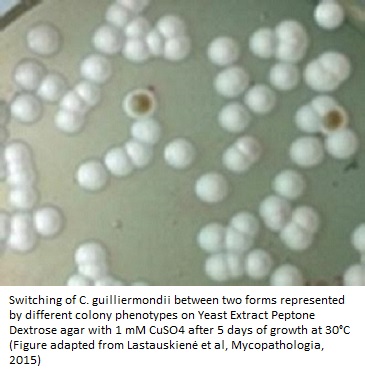
A group of scientists from the Faculty of Natural Sciences, Vilnius University, and State Research Institute Centre for Innovative Medicine, Lithuania published an article in Mycopathologia in 2015 (3). This group investigated for the first time whether the phenotypic switching (switching of fungal microbes between different cellular types) occurs in C. guilliermondii; and whether this switching could attribute to a change in drug susceptibility of the pathogen.
Phenotypic switching generally allows the pathogen to attain different morphologies depending upon the surroundings and living conditions in the environment. The yeast form carries out the role of dissemination while the hyphae executes the adhesion to host cell, tissue invasion, and proteolytic activity using enzymes like lipases and proteases. Thus, they can easily adapt to stringent conditions (high temperatures, wide range of pH, and nutrient starvation stress) and invade the host cells, leading to destruction and increased virulence.
In the aforementioned studies, the switch was seen in the form of white or brown colonies on YPD agar and displayed a much higher resistance to the antifungal amphotericin B (nearly 20 times!) and formic acid (3).
On the other hand, if the fungi do not show major phenotypic switching, it is generally associated with reduced virulence or no virulence at all. Phenotypic switching is already known in case of C. albicans, C. dubliniensis, and C. tropicalis and is controlled by genes encoded by the MTL (mating-type-like) loci. In addition, the pathogenic C. glabrata exhibits core switching between white, light brown, dark brown, very dark brown, and irregular wrinkle colonies. Another species, C. lusitaniae that shows antifungal resistance, undergoes phenotypic switching between white, light brown, and dark brown colonies similar to C. glabrata (4, 5).
C. guilliermondii has been isolated from a number of sources, both human infection sites, as well as environmental samples. These include human skin and mucosal surfaces, soil, insects, plants, sea water and secretions produced by various trees (6). This yeast-like species is known to produce only pseudohyphae and grows only on Sabouraud dextrose agar (SDA). It shows small, flat, or smooth colonies (yellowish or creamish) on the agar along with short pseudohyphae. P. guilliermondii and P. caribbica can grow and assimilate D-xylose, ribose, ramnose, melibiose, lactose, sorbose, and esculine and do not assimilate lactate (6).
How is C. guilliermondii Different From C. albicans and Who are Susceptible?
As far as the intensity of infection is concerned, C. guilliermondii is comparatively less virulent than C. albicans, though it has exhibited higher resistance to antifungal treatments.
C. guilliermondii is associated with a number of co-morbidities and accompanying risk factors including presence of solid tumors and blood-associated malignancies (acute non-lymphoid leukemia, acute lymphoid leukemia, non-Hodgkin’s lymphoma, multiple myeloma); patients undergoing organ and bone marrow transplantations (exhibiting Graft-versus-host disease); prolonged treatments to antibiotics, steroids, long-term antifungals (mostly fluconazole, amphotericin B, nystatin, caspofungin), chemotherapy, bacteremia; neutropenia; patients having implanted medical devices such as intravascular catheters, and dental devices.
Moreover, individuals who are, in general, hospitalized, admitted to ICUs, and have undergone surgeries are vulnerable to developing C. guilliermondii infections. Since it also has to do with dysbiosis and imbalance of normal microflora in human gut, individuals complaining of inflammatory bowel disease and gastric ulcers may also show the symptoms of C. guilliermondii fungal infections (1, 7).
Implications of Candida guilliermondii Infections
C. guilliermondii is an uncommon fungal yeast pathogen which behaves as a human saprophyte. It causes serious infections in immunocompromised hosts, especially patients with hematological malignancies (cancer of blood). A number of clinical symptoms and syndromes can be attributed to the C. guilliermondii infections. These include white or yellow discoloration of nails (onychomycosis), infection in the endocardial lining of internal chambers of heart (endocarditis due to spreading of fungi/bacteria from other parts of the body into heart via blood) hospital acquired nosocomial candidemia, infections in the bones (osteomyelitis), septic arthritis, and candiduria i.e., mixed yeast infections in urine, probably due to bladder colonization (1, 7-9).
A study carried out at Santa Casa Complexo Hospitalar, Brazil in 11 patients with C. guilliermondii infection showed that immunocompromised host immunity (solid organ transplantation, AIDS, and leukemia) was a major factor in such infections. These patients had undergone procedures like central venous catheter insertions, abdominal surgery, and peritoneal dialysis. While the risk factors for candidemia in these patients were similar to those of other Candida yeast species, the mortality associated with C. guilliermondii was significantly lower (10). This study also reflects the general etiology of C. guilliermondii infections to range from 0.1% to 2.8% among approximately 850 samples tested positive for Candida species in the Brazilian medical settings.
Another study conducted at Madrid, Spain (published in Antimicrobial Agents and Chemotherapy in 2017) performed the microbial characterization of C. guilliermondii isolates and compared them with C. albicans in terms of biofilm formation, antifungal resistance, and metabolic activities. Scanning electron microscopy revealed that the C. guilliermondii complex biofilms comprised primarily of clamped blastospore layers lacking hyphae or pseudohyphae (2, 11).
Yet another study carried out collaboratively between Università La Sapienza and Pfizer Italia, Rome also reviewed the infections in patients with hematological malignancies caused by C. guilliermondii in their department. This study recorded an occurrence of 12% cases of C. guilliermondii infections among the total cases of candidemia. The major contributors to this 12% were use of central venous catheters and invasive tissue infections. The C. guilliermondii clinical isolates were susceptible amphotericin B, voriconazole, and fluconazole and comparatively exhibited greater resistance to flucytosine, itraconazole, and caspofungin in this study (12).
Antifungal Resistance and Treatment
One of the major concerns of emergence of all these yeast species is resistance to the available antifungal drugs. Antifungal resistance is characterized by increased minimum inhibitory concentrations, poor treatment results, and increase in the spread of invasive infections. This can be ascribed to a number of mechanisms which can enable these Candida species to acquire resistance. One of these mechanisms is overexpression of the efflux pump encoding genes like MDR or CDR, and occurrence of point mutations in the ERG11 enzyme coding genes. To quote an example, point mutations in the FKS genes in the genomes of a few of the Candida species confer echinocandin resistance (5).
An Italian hospital has also reported a nosocomial cluster of C. guilliermondii fungemia in five clinical isolates that were resistant to 5-flucytosine and responded well to fluconazole and amphotericin B (13). The damage due to candidemia could be undone only upon the removal of vascular catheters and fluconazole treatment. Though fluconazole may be effective most of the time, there are also cases of acute resistance against this drug in a number of clinical C. guilliermondii infections.
In an extreme case, C. guilliermondii infection of the bone in one of the fingers of the patient (osteomyelitis) left the patient with no other choice than amputation of that finger. The C. guilliermondii strain isolated from this amputated digit was found to be resistant to both fluconazole and itraconazole (14).
A study at the National Taiwan University Hospital showed that among C. guilliermondii isolates, 98%, 100%, and 98% showed sensitivity to caspofungin, micafungin, and anidulafungin, respectively (15). Most of the studies believe that voriconazole therapy is more efficient in treating C. guilliermondii than fluconazole, and the resistance rate towards voriconazole is also low comparatively.
To learn how to treat Candida guilliermondii naturally click here.
About the Author

Dr. Vibhuti Rana completed her Bachelors's Degree (Bioinformatics
Hons.) from Punjab University and accomplished her Master’s Degree
(2012) in Genomics with a Gold Medal from Madurai Kamaraj University,
India. In 2020, she received her doctorate in Molecular Biology from the
Council of Scientific and Industrial Research-Institute of Microbial
Technology in affiliation with the Jawaharlal Nehru University, New
Delhi, India.
Her focus areas include microbial drug resistance, epidemiology, and protein-protein interactions in infectious diseases. As a Molecular Biologist with extensive experience with infectious diseases, we are happy she is part of the YeastInfectionAdvisor team.
If you have any questions about Candida guilliermondii or yeast infections in general, please feel free to contact us through the contact page on this website or see your doctor.
Dr. Rana's Medical References
- Savini V, Catavitello C, Onofrillo D, et al. What do we know about Candida guilliermondii? A voyage throughout past and current literature about this emerging yeast. Mycoses. 2011; 54(5):434-441. doi:10.1111/j.1439-0507.2010.01960.x. https://pubmed.ncbi.nlm.nih.gov/21039941/
- Castillo-Bejarano JI, Tamez-Rivera O, Mirabal-García M et al. Invasive Candidiasis Due to Candida guilliermondii Complex: Epidemiology and Antifungal Susceptibility Testing From a Third-Level Pediatric Center in Mexico, Journal of the Pediatric Infectious Diseases Society, 9 (3), 2020, Pages 404–406, https://doi.org/10.1093/jpids/piaa043. https://academic.oup.com/jpids/article-abstract/9/3/404/5856407
- Lastauskienė E, Čeputytė J, Girkontaitė I, Zinkevičienė A. Phenotypic switching of Candida guilliermondii is associated with pseudohyphae formation and antifungal resistance. Mycopathologia. 2015;179(3-4):205-211. doi:10.1007/s11046-014-9844-3. https://pubmed.ncbi.nlm.nih.gov/25481846/
- Lachke SA, Joly S, Daniels K, Soll DR. Phenotypic switching and filamentation in Candida glabrata. Microbiology. 2002;148(Pt 9):2661-2674. doi:10.1099/00221287-148-9-2661. https://pubmed.ncbi.nlm.nih.gov/12213913/
- Pfaller MA. Antifungal drug resistance: mechanisms, epidemiology, and consequences for treatment. Am J Med. 2012; 125(1 Suppl):S3-S13. doi:10.1016/j.amjmed.2011.11.001. https://pubmed.ncbi.nlm.nih.gov/22196207/
- Desnos-Ollivier M, Ragon M, Robert V, Raoux D, Gantier JC, Dromer F. Debaryomyces hansenii (Candida famata), a rare human fungal pathogen often misidentified as Pichia guilliermondii (Candida guilliermondii). J Clin Microbiol. 2008; 46(10):3237-3242. doi:10.1128/JCM.01451-08. https://pubmed.ncbi.nlm.nih.gov/18701668/
- Tseng TY, Chen TC, Ho CM, et al. Clinical features, antifungal susceptibility, and outcome of Candida guilliermondii fungemia: An experience in a tertiary hospital in mid-Taiwan. J Microbiol Immunol Infect. 2018;51(4):552-558. doi:10.1016/j.jmii.2016.08.015. https://pubmed.ncbi.nlm.nih.gov/28625801/
- Pfaller MA, Diekema DJ, Mendez M, et al. Candida guilliermondii, an opportunistic fungal pathogen with decreased susceptibility to fluconazole: geographic and temporal trends from the ARTEMIS DISK antifungal surveillance program. J Clin Microbiol. 2006; 44(10):3551-3556. doi:10.1128/JCM.00865-06. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1594787/
- Borg-von Zepelin M, Kunz L, Rüchel R, Reichard U, Weig M, Gross U. Epidemiology and antifungal susceptibilities of Candida spp. to six antifungal agents: results from a surveillance study on fungaemia in Germany from July 2004 to August 2005. J Antimicrob Chemother. 2007;60(2):424-428. doi:10.1093/jac/dkm145. https://pubmed.ncbi.nlm.nih.gov/17562683/
- Pasqualotto, Alessandro Comarú, Antunes, Ana Graciela Ventura, & Severo, Luiz Carlos. (2006). Candida guilliermondii as the aetiology of candidosis. Revista do Instituto de Medicina Tropical de São Paulo, 48(3), 123-127. https://doi.org/10.1590/S0036-46652006000300002. https://www.scielo.br/scielo.php?script=sci_arttext&pid=S0036-46652006000300002
- Marcos-Zambrano LJ, Puig-Asensio M, Pérez-García F, et al. Candida guilliermondii Complex Is Characterized by High Antifungal Resistance but Low Mortality in 22 Cases of Candidemia. Antimicrob Agents Chemother. 2017; 61(7):e00099-17. Published 2017 Jun 27. doi:10.1128/AAC.00099-17. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5487632/
- Girmenia C, Pizzarelli G, Cristini F, et al. Candida guilliermondii fungemia in patients with hematologic malignancies. J Clin Microbiol. 2006; 44(7):2458-2464. doi:10.1128/JCM.00356-06. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1489483/
- Masala L, Luzzati R, Maccacaro L, Antozzi L, Concia E, Fontana R. Nosocomial cluster of Candida guillermondii fungemia in surgical patients. Eur J Clin Microbiol Infect Dis. 2003; 22(11):686-688. doi:10.1007/s10096-003-1013-4. https://pubmed.ncbi.nlm.nih.gov/14566575/
- Tietz HJ, Czaika V, Sterry W. Case report. Osteomyelitis caused by high resistant Candida guilliermondii. Mycoses. 1999; 42(9-10):577-580. doi:10.1046/j.1439-05 07.1999.00497.x. https://pubmed.ncbi.nlm.nih.gov/10592705/
- Chen CY, Huang SY, Tang JL, et al. Clinical features of patients with infections caused by Candida guilliermondii and Candida fermentati and antifungal susceptibility of the isolates at a medical centre in Taiwan, 2001-10. J Antimicrob Chemother. 2013; 68(11):2632-2635. doi:10.1093/jac/dkt214. https://pubmed.ncbi.nlm.nih.gov/23766486/
Home Privacy Policy Copyright Policy Disclosure Policy Doctors Store
Copyright © 2003 - 2025. All Rights Reserved under USC Title 17. Do not copy
content from the pages of this website without our expressed written consent.
To do so is Plagiarism, Not Fair Use, is Illegal, and a violation of the
The Digital Millennium Copyright Act of 1998.